María José Gosalbes, Carlos David Esteban, José Luis Galán and Gaspar Pérez-Martínez.
Published in Applied and Environomental Microbiology (2000) 66 (11): 4822-4828
Pubmed link: http://www.ncbi.nlm.nih.gov/pubmed/11055930. Full text:
ABSTRACT :
Lactose operon from Lactobacillus casei is regulated by a very tight glucose repression and substrate induction mechanisms, which made it a tempting candidate system for the expression of foreign genes or metabolic engineering. An integrative vector was constructed allowing stable gene insertion in the chromosomal lactose operon of Lactobacillus casei. This vector is based on the non-replicative plasmid pRV300 containing two DNA fragments corresponding to the 3’ end of lacG and the complete lacF gene. Four unique restriction sites were created, as well as a ribosome binding site that would allow the cloning and expression of new genes between these two fragments. Then, integration of the cloned genes into the lactose operon of L. casei could be achieved via homologous recombination in a process that involved two selection steps, which yielded highly stable food-grade mutants. This procedure has been successfully used for the expression of E. coli gusA and L. lactis ilvBN genes in L. casei. Following the same expression pattern as the lactose genes, b-glucuronidase activity and diacetyl production were repressed by glucose and induced by lactose. This integrative vector represents a useful tool for strain improvement in Lactobacillus casei, that could be applied to engineering fermentation processes or for expression of genes for clinical and veterinary uses.
INTRODUCTION:
Lactic acid bacteria (LAB) have been used for centuries in the preparation and processing of foods and beverages. Due to its great economic importance for the agro-feed sector and its alleged importance for human and animal health, research on characterization, metabolism and genetics of the genus Lactobacillus has increased over the last decade. Several vectors have been developed to express genes and to secrete proteins in Lactobacillus (12,26,36,37,47,48). However, if they are to be considered safe for humans, animals or the environment, only DNA from GRAS organisms (generally regarded as safe) and no antibiotic resistance markers should remain after genetic manipulation. The integration of foreign genes in the genome constitutes an interesting option to stably maintain cloned genes without the need for selective markers. Technically, foreign gene integration could be achieved by homologous recombination through cloned DNA fragments (randomly cloned fragments or target genes) and by self-integrative elements (insertion sequences or attachment site and integrase gene). In the genus Lactobacillus, stabilization of cloned genes is normally achieved by chromosomal integration, based on the use of cloned DNA fragments in non-replicating plasmids. Stable chromosomal integration of the genes encoding the a-amylase from Bacillus stearothermophilus and a cellulase from Clostridium thermocellum was obtained in Lactobacillus plantarum using a randomly cloned chromosomal fragment as the integration target (42). A similar strategy was used to construct an integrative vector for Lactobacillus acidophilus (25). Other ingenious systems have been developed using a phage integrase-mediated site-specific insertion in the host chromosome (4,6,28). There are also examples of stable integration in target genes, such as cbh, that encodes a bile salt hydrolase and pepXP, encoding an X-prolyl-dipeptidyl aminopeptidase from L. plantarum and L. helveticus, respectively (9,22). All foreign genes integrated by these procedures are normally expressed from their own promoters which makes more difficult the control of their regulation. The a-amylase gene from Bacillus licheniformis (amyL) was satisfactorily expressed in L. plantarum only when the amyL promoter was replaced by a L. plantarum promoter (42).
Very efficient expression systems based on antimicrobial peptide (nisin), sugar utilization or nonsense suppressors have been developed in Lactococcus lactis (12,14,23,40). However, besides the nisin system these approaches could not be transferred to species of Lactobacillus. In lactobacilli, the regulation of gene expression has been mainly studied for carbon catabolism pathways, such as those of lactose, xylose, ribose, sorbose and arginine deiminase (1,2,3,10,18,19,30,34,35,43,49,50). In Lactobacillus casei, the best-characterized sugar transport is the lactose-specific phosphoenolpyruvate-dependent phosphotransferase system (PTS). The lac operon, lacTEGF, encodes an antiterminator protein (LacT), lactose-specific PTS proteins (LacE and LacF) and a phospho-b-galactosidase (LacG) (1,2,3,18,35). It has been previously reported (3,19,30) that the expression of the lac operon in L. casei ATCC393 [pLZ15–] is subject to dual regulation: carbon catabolite repression (CCR) mediated by the general regulator CcpA and induction by lactose through transcriptional antitermination. LacT whose activity is modulated by the EII elements of the lactose PTS (LacE and/or LacF) mediates this later mechanism. Furthermore, HPr, a general component of PTS, and LacT are involved in an additional CcpA-independent CCR effect (19,39,44).
In this report, an integrative expression vector is described that allowed the selection of stable mutants that express E. coli gusA and L. lactis ilvBN genes through the lactose regulon. This integrative vector represents the first specific expression system developed for L. casei with a great potencial in food industry and health applications.
MATERIALS AND METHODS:
Bacterial strains, plasmids and growth conditions.
The strains and plasmids used in this work are listed in Table 1. L. casei cells were grown in MRS medium (Oxoid) and MRS fermentation broth (Adsa-Micro, Scharlau S.A., Barcelona, Spain) plus 0.5% of the different carbohydrates at 37°C under static conditions. E. coli DH5a was grown with shaking at 37°C in Luria-Bertani (LB) medium. Plating of bacteria was performed on the respective media solidified with 1.5% agar. When required, the concentrations of antibiotics used were 100 mg of ampicillin per ml to select E. coli transformants and 5 mg of erythromycin per ml for L. casei.
TABLE 1: Strains and plasmids used in this study.
Strain or plasmid | Description | Source or reference |
| Strains | ||
| Lactobacillus casei | CECTa 5275 | B. Chassy (Univ. Illinois, Urbana, IL61801) |
| L. casei CECT 5276 | L. casei CECT 5275with a frame shift in lacF | (19) |
| L. casei CECT 5290 | L. casei CECT 5275 with gusA gene integrated | This work |
| L. casei CECT 5291 | L. casei CECT 5275 with ilvBN genes integrated | This work |
| Lactococcus lactis MG1363 | MG1363 | (16) |
| Plasmids | ||
| pRV300 | Erir from pAMb1 | (24) |
| pNZ272 | gusA Cmr | (33) |
| PIlacF | lacF gene cloned as EcoRV/KpnI fragment in pRV300 | This work |
| pIlac | lacG 3’end and lacF gene in pRV300 | This work |
| pIlacgus | PIlac containing gusA gene | This work |
| pIlacilv | PIlac containing ilvBN genes | This work |
(a) CETC: Colección Española de Cultivos Tipo
Recombinant DNA procedures.
Genomic DNA from Lactobacillus casei and Lactococcus lactis strains was purified using Puregene DNA isolation Kit (Gentra Systems, Inc., Minneapolis, USA), following the procedure described by the manufacturer. Restriction and modifying enzymes were used according to the recommendations of manufacturers. General cloning procedures were performed as described by Sambrook et al. (41). L. casei was transformed by electroporation with a Gene-pulser apparatus (Bio-Rad Laboratories, Richmond, Calif.). For Southern blot hybridization, L. casei DNA was digested with BglII and PstI endonucleases, separated on agarose gel and blotted to Hybond nylon membrane (Amersham). The probes used in the three hybridization experiments were Perm, Pilv and Plac. Perm probe consists in erm gene from pRV300 plasmid digested with BamHI endonuclease. Pilv corresponds to ilvBN genes from L. lactis that was obtained by PCR using ilv1 and ilv2 oligonucleotides as primers (sequence see below) and the genomic DNA of Lactococcus strain as template. Plac probe comprised a fragment of 537 base pairs (bp) that corresponds to the 3’ end of lacG gene. This DNA fragment was obtained by PCR using genomic DNA from L. casei as template and the oligonucleotides lac46 (5’TGCGTGCCTATCATGGC) and lac6 (5’CTTGCTGTCTAAATAGCC) as primers. The DNA probes were prepared using the reagents from the Boehringer digoxigenin-DNA labelling kit as recommended by the manufacturer. Hybridization, washing and staining were done as described by the supplier. PCR was performed using Expand TM High fidelity PCR system (Roche Molecular Biochemicals), containing 200 mM of each deoxynucleoside triphosphate and 10 pmol of each primer. Upon agarose gel electrophoresis, the amplified DNA was recovered with GFX TM PCR Kit (Amersham Pharmacia Biotech).
Construction of integration plasmids.
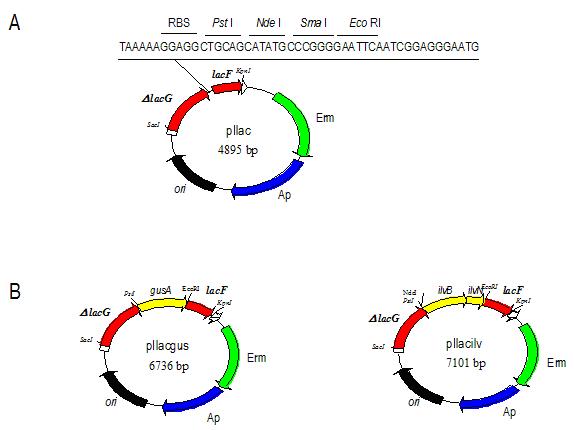
The integrative vector, pIlac, is based on vector pRV300 (24) that is not replicating in Lactobacillus and it carries the erm gene from pAMb1. This vector was constructed in a two steps cloning experiment. A 461 bp DNA fragment, containing lacF gene, was amplified by PCR from L. casei chromosome with the primers lac43 (5’TACATATGCCCGGGGAATTCAATCGG AGGGAAAATG) and lac45 (5’TTGAGGTACCGCTAACAGC). Lac43 primer showed several substitutions (boldface) to introduce the new NdeI, SmaI and EcoRI sites (underlined) in the 45 bp region between lacG and lacF. Lac45 contained three substitutions (boldface) generating a new KpnI site (underlined). The blunt-ended fragment amplified was digested with KpnI and cloned into pRV300 previously digested with EcoRV and KpnI. This construction, pIlacF, was used to clone an 878 bp DNA fragment, containing the 3’end of lacG gene and 23 bp of the 45 bp intergenic region between lacG and lacF genes. This fragment was amplified by PCR from L. casei chromosome using the primers lac49 (5’ATAAGAGCTCCCAAGCTGA) and lac42 (5´TGCATATGCTGCAGCCTCCTTTTTAATCCGGAATG). Lac49 had three substitutions (boldface) generating a SacI site (underlined). Lac42 primer contained several substitutions (boldface) creating new NdeI and PstI sites and a ribosome-binding site (RBS) (underlined). Then, this fragment was cloned in the NdeI/SacI sites of pIlacF. Figure 1A shows the physical map of the resulting integrative vector, pIlac.
A DNA fragment, containing E. coli gusA gene, was amplified by PCR using the plasmid pNZ272 (33) as template and the primers gus1 (5’AAAACTGCAGTATTATTATCTTAATGAGG) (a newly created PstI site is underlined) and gus2 (5’CGGAATTCTCATTGTTTGCCTCCC) (a newly created EcoRI site is underlined). The amplified DNA fragment was purified, digested with the endonucleases EcoRI and PstI and ligated into an EcoRI/PstI digested pIlac. This construction was named pIlacgus (Figure 1B).
The ilvBN genes were amplified from the genomic DNA of L. lactis using the oligonucleotides ilv1 (5’CGATCATATGAAAAAAATAAAGTTAGAAAAACCTA CTTCC) and ilv2 (5’CCGAATTCTTAGCCACGCTCAAAACCTGC) as primers, containing NdeI and EcoRI sites (underlined), respectively. The amplified fragment was cloned into NdeI/EcoRI digested pIlac to give pIlacilv (Figure 1B).
Enzymatic assays.
Phospho-b-galactosidase (P-b-gal) and b-glucuronidase activities were assayed as previously described (33,46) in permeabilized L. casei cells.
Total nitrogen determination.
The procedure used was based on the method described by Doi et al. (15).
Determination of end metabolites.
The metabolites released by wild-type and mutant strains grown on glucose + lactose, lactose or ribose have been analyzed in a resting cell system (11). The determination of volatile compounds such as ethanol, acetaldehyde, acetone, 1-butanol, acetoin and diacetyl was performed using a purge-and-trap (PAT) apparatus equipped with a Vocarb 3000 trap (Supelco) to concentrate the analytes and coupled to a gas chromatographer equipped with a mass spectrometer (Hewlett-Packard 7695, Barcelona, Spain) as described Dauneau et al. (11). The a-acetolactate (ALA) determination is based on its oxidative descarboxylation to diacetyl as previously reported with some modification (38). The samples (2 ml) were pretreated in a 4 ml vial by addition of 150 µl of 1.85 M FeCl3 and 1 ml of 80% lactate buffer (pH 2.8) and vigorously stirred for 5 seconds. The vials were hermetically sealed with teflon-lined rubber seals and heated at 75ºC for 30 min. The concentration of ALA was calculated by subtraction of the diacetyl concentration found before and after decarboxylation of the samples.
Lactic acid produced in the resting cell system was measured with a D-lactic acid/L-lactic acid enzymatic bioanalysis kit (Boehringer-Mannheim) as described by the supplier. The total amount of lactic acid produced corresponds to the addition of the concentration of both isomers determined.
RESULTS:
Integration strategy with vector pIlac.
This vector (Fig.1A) contains two regions of homology that are physically close in L. casei chromosome, a 3’ fragment of lacG (D lacG) and lacF. The sequence of the intergenic region has been modified, remaining intact the spacing between both genes, to introduce a typical RBS for Lactobacillus and a multiple cloning site (PtsI, NdeI, SmaI and EcoRI), allowing the cloning of new genes. So that, after integration the transcription of these genes would take place from the lac promoter and the newly created RBS would facilitate their translation initiation.
For the integration of the cloned genes in the chromosome, two recombination events should take place, one in each of the homologous regions (DlacG and lacF). The use of a frame-shift mutant in lacF (19), L. casei CECT 5276, as host strain (Lac–) facilitated the selection of the clones that had undertaken the second recombination as Lac+ clones among the Lac– background, with the following procedure. After electroporation of L. casei CECT 5276 with pIlac derivatives, Lac+Er+ and Lac–Er+ transformants were recovered, depending on the region where the Campbell-like recombination had occurred. A transformant with Lac–Err phenotype was grown for 200 generations in MRS broth without antibiotic in order to allow the second recombination event, which would excise the plasmid rendering Lac+Ers colonies (one out of 20 viables). This strategy can be applied for the insertion of any gene of interest in the lac operon of L. casei and rendered mutants totally deprived of any DNA sequence from the plasmid and erm gene.
Chromosomal integration of E. coli gusA in L. casei.
In order to evaluate the potencial of the integrative vector pIlac as vehicle for chromosomal gene insertion, the b-glucuronidase encoding gene of E. coli gusA was cloned in it. The plasmid obtained, pIlacgus (Fig.1B), was used to transform L. casei CECT 5276. Following the procedure described above, colonies that had undergone a second recombination event suffered the excision of the vector, giving rise to Erms Lac+ colonies which had integrated the gusA gene in the lac operon. First and second recombination events were confirmed by Southern blot hybridization of the integrants chromosomal DNA (data not shown). The resulting new structure of the lac operon contained gusA between lacG and lacF, and as consequence, the expression of gusA was subject to the same regulation as the lac genes. This was confirmed by measuring b-glucuronidase activity in one of the colonies selected, L. casei CECT 5290, when grown on ribose, lactose and glucose+lactose (Table 2). Greater P-b-galactosidase activity was detected in the gusA integrant on lactose than the wild-type and ilvBN integrant (described below), possibly due to the partial cleavage of ONPG-6-P (P-b-gal substrate) by b-glucuronidase. It could also be noticed that the growth rate of both integrants on glucose was identical to the wild type (data not shown). On lactose, duplication times were not substantially different during early growth stages (95.5 ± 2.6 min; 91.8 ± 3.4 min; 90.3 ± 5.4 min for wild-type, CECT 5290 and CECT5291 strains, respectively), however, it was observed that both mutants would only reach OD550 = 0.8 .
Construction of a food-grade ilvBN integrant of Lactobacillus casei.
Diacetyl is an important compound related to the characteristic flavor of many fermented milk products. Only a few LAB could produce this metabolite from citrate of the milk. During citrate fermentation, a-acetolatate synthase converts pyruvate to a-acetolatate that could be converted spontaneously to diacetyl in the presence of oxygen. Lactococcus lactis ilvBN genes encode the catalytic and regulatory subunits of acetohydroxy acid synthase, respectively (17). This enzyme is involved in biosynthesis of branched-chain amino acids, isoleucine and valine, converting pyruvate to a-acetolactate with higher affinity for pyruvate than a-acetolatate synthase. In this work, in order to increase the cellular pool of a-acetolactate that could be turned into diacetyl by oxidative decarboxylation, ilvBN genes of L. lactis were integrated in the chromosome of L. casei. Both genes were cloned into pIlac to give pIlacilv (Fig.1B) which was used to transform L. casei CECT 5276. The selection strategy of recombinant colonies was identical to that described above. A double recombinant (Erms and lactose+) of L. casei (ivlBN integrant) was selected for further analysis and designated as L. casei CECT 5291. The pattern of P-b-gal activity in this strain was similar to that in wild-type strain, as it is also induced by lactose and repressed by glucose (Table 2).
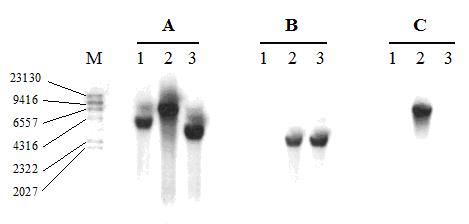
Integration of ilvBN genes into the chromosomal lac operon of L. casei was confirmed by Southern hybridization using the probes Plac, corresponding to the 3´end of lacG (Figure 2A). A hybridization band was detected on the genomic DNA of L. casei CECT 5276 (host strain), Ermr lactose– integrant of pIlacilv (first recombination event) and L. casei CECT 5291 (double recombinant) (Fig.2A lanes 1, 2 and 3, respectively) when digested with BglII/PstI. The fragment detected in L. casei CECT 5276 was larger than in the double recombinant because the fragment integrated containing ilvBN carries an additional PstI site. The chromosomal integration of ilvBN was confirmed using Pilv probe (Figure 2B). Evidence that the antibiotic resistance gene (erm) had been excised from L. casei genome was demonstrated using Perm probe. In this Southern blot hybridization signal was only detected with the genomic DNA from the first integrant (Figure 2C). Moreover, the ilvBN genes remained stably integrated on the genome after 50 overnight transfers in MRS medium without selective pressure (data not shown).
TABLE 2: Enzymatic activities in different strains of L. casei.
Strain | Relevant genotype | Sugar | P-b-galactosidase (nmol min-1mg-1 dry wt) | b-glucuronidase (nmol min-1mg-1 dry wt) |
| Ribose | 2.03 ± 0.86 | 0.98 ± 0.15 | ||
| L. caseiCECT 5275 | wild-type | Lactose | 23.91 ± 2.93 | 1.89 ± 0.19 |
| Glucose + Lactose | 0.78 ± 0.24 | 1.16 ± 0.23 | ||
| ||||
| Ribose | 10.040 ± 1.69 | 43.13 ± 11.64 | |
| L. caseiCECT 5290 | gusA | Lactose | 57.84 ± 8.51 | 119.05 ± 8.91 |
| Glucose + Lactose | 1.26 ± 0.12 | 1.23 ± 0.2 | |
| Ribose | 6.43 ± 0.63 | ||
| L. caseiCECT 5291 | ilvBN | Lactose | 27.11 ± 3.56 | |
| Glucose + Lactose | 1.56 ± 0.26 |
The values and standard deviations are at least from three independent experiments.
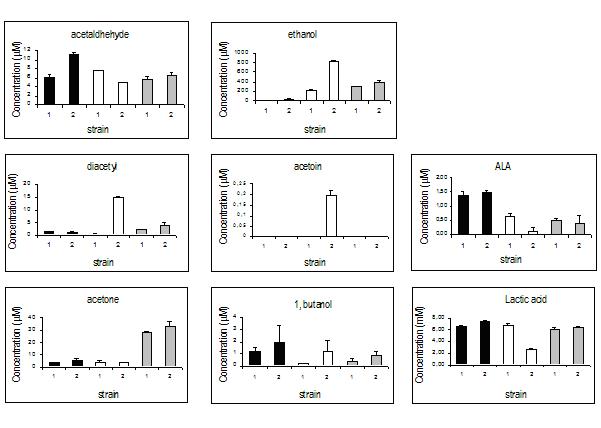
Determination of metabolic products in the ilvBN mutant.
Metabolites released by L. casei wild-type and L. casei CETC 5291, carrying the integrated ilvBN genes in the lactose operon (ilvBN integrant), have been analyzed in a resting cell system when cells were grown on glucose+lactose, lactose and ribose (Fig. 3). Besides lactic acid, which is by far the most abundant compound, ethanol and acetone were the predominant metabolites accumulated by ribose-grown cells, where L. casei becomes heterofermentative. Remarkable differences could be noticed in the ilvBN integrant on lactose, regarding the production of ethanol, 1-butanol, acetoin and diacetyl. In particular, the amount of diacetyl accumulated by the lactose-induced ilvBN integrant was 23 fold greater than that of the wild type. Other significant differences found in the ilvBN integrant are related to the lower accumulation of ALA and lactate on lactose, possibly due to the diversion of pyruvate and ALA towards the synthesis of branched-chain aminoacids (isoleucine, leucine, valine). In order to test this hypothesis, total soluble nitrogen (aminoacids) was determined in the supernatant of the resting cell systems on lactose, obtaining 1.25 ± 0.07 mM, in the wild type, and 2.01 ± 0.26 mM, in the ilvBN integrant. This difference (0.76 mM) could partially be explained by the secretion of a proportion of the excess aminoacids synthesized by the integrant. Regarding acetaldehyde production, only small differences were observed between both strains with different carbon sources. Unexpectedly, a slightly higher concentration of acetaldehyde and also 1-butanol could be observed when the ilvBN integrant was grown on glucose+lactose. This different behavior was clearly related to the presence of the ilvBN genes indicating that some degree of expression of the acetohydroxy acid synthase was taking place on glucose+lactose, altering the proportions in metabolites derived from acetyl-CoA.
DISCUSSION:
The present study describes the construction of an integrative expression vector for Lactobacillus casei that allowed obtaining stable food-grade integrants capable to express foreign genes under the tight control of the well-characterized lac operon promoter (Fig.1A). Lactose genes have been used in other LAB for different biotechnological purposes, such as the construction of food-grade vectors in Lactococcus lactis, to address integration in Lactobacillus helveticus and for gene expression in Streptococcus thermophilus (9,27,29,31). However, both, structural organization and regulation of the lac operon in L. casei are very different to those described in the other LAB (1,2,3,13,18,19,30,35); it displays very tight glucose repression and lactose induction mechanisms, which were very promising for the expression of foreign genes. The integrative vector designed in this work, pIlac, allowed cloning of DNA fragments between the two target genes (DlacG and lacF). Then, through Campbell-like recombination, the genes of interest could be inserted in the lac operon, obtaining a food-grade construct in which the foreign genes became a functional part of the operon and were subject to the same regulation (Table 2). However, it could be observed that the insertion of foreign genes, gusA and ilvBN, lead to some induction of the operon on ribose, as shown by P-b-gal and b-glucuronidase activities (Table 2). In a previous work it was shown that mutants in the genes encoding either of the lactose-specific PTS elements (lacE, lacF) showed constitutive transcription of the lac operon (19). It could be tentatively proposed that transcription of the new constructions gave a longer, more unstable mRNA, where perhaps lacF -placed at the end of it- could be less efficiently translated. This could be confirmed by the fact that neither of the integrants could grow on lactose more than a few generations, possibly because of an inefficient lactose-PTS transport that allowed growth only at high lactose concentrations.
L. casei is frequently used as starter culture in many fermentation processes, especially in cheese making or recently, as probiotic in fermented milk products. However, L. casei is not a good diacetyl producer and this is a very desirable compound in dairy fermentations. This product is normally synthesized in LAB from the glycolytic intermediate pyruvate, which is converted to a-acetolactate (ALA) by the a-acetolactate synthase. Then, ALA is transformed to acetoin through a-acetolactate decarboxylase activity or to diacetyl in presence of oxygen. Also, acetoin yields diacetyl by action of acetoin reductase. Different approaches have been used in L. lactis to improve diacetyl production, such as deletion of acetolactate decarboxylase, mutation of lactate dehydrogenase or overexpression of acetohydroxy acid synthase (8,20,21,32,45). This biotechnological approach was also considered in the present work. Induction of ilvBN genes by lactose in the food-grade system developed in this work yielded, in three hours, a total amount of diacetyl comparable to that in overnight cultures of L. lactis overexpresing ilvBN (7,8,21,45). However, further optimization of the diacetyl production through detailed fermentation studies could be achieved because, in our resting cell system, the high cell density and static incubation conditions in a sealed tube was possibly generating an adverse environment -poor in oxygen -, where a lower diacetyl reductase activity favored certain accumulation of acetoin.
Another objective of this work was the evaluation of the metabolites balance during the overexpression of “cross-road” enzymes such as acetohydroxy acid synthase (encoded by ilvBN). The accumulation of ALA on glucose-grown cells suggests that the biosynthesis of diacetyl from this ketoacid could be subject to CCR. However, a major interference with an even greater overproduction of diacetyl in L. casei CETC 5291 can be attributed to the fact that pyruvate is also substrate of acetohydroxy acid synthase in the anabolic pathway of branched-chain aminoacids. Furthermore, both compounds, pyruvate and ALA, could be inducers of subsequent steps in these pathways (5), for which the overexpression of ilvBN was probably leading to a drainage of pyruvate for the synthesis of leucine, isoleucine and valine. This was demonstrated by the lower amount of lactate and higher concentration of aminoacids detected in the supernatant of the lactose-induced mutant during the resting cell assay. However, a metabolite balance could not be calculated. This is an intricate part of the metabolic map and many more compounds should be determined to get a clearer picture of the carbon fluxes at this metabolic level.
The kind of experiments described in this work has never before been performed in lactobacilli. Due to its food-grade nature, the system developed here has a great potential for the metabolic engineering of intracellular metabolites and the production of different enzymes during dairy fermentation. However, due to its regular presence in higher vertebrate mucosae, other applications could be envisaged, such as the delivery of antigens in the gut, mouth or vagina or other clinical and veterinary products.
ACKNOWLEDGMENTS:
We are thankful to M.C. Miralles for her skillful technical assistance. This work was financed by the EU project BIO4-CT96-0380 and by founds of the Spanish CICyT (Interministerial Commision for Science and Technology) (Ref. ALI 98-0714). C.D.E. was the recipient of a fellowship of Spanish government.
REFERENCES:
- Alpert C.-A., and B. M. Chassy. 1988. Molecular cloning and nucleotide sequence of the factor IIIlac gene of Lactobacillus casei. Gene. 62:277-288.
- Alpert C.-A., and B. M. Chassy. 1990. Molecular cloning and DNA sequence of lacE, the gene encoding the lactose-specific enzyme II of the phosphotransferase system of Lactobacillus casei. J. Biol. Chem. 265:22561-22568.
- Alpert, C.-A., and U. Siebers. 1997. The lac operon of Lactobacillus casei contains lacT, a gene coding for a protein of BglG family of transcriptional antiterminators. J. Bacteriol. 179:1555-1562.
- Alvarez, M. A., M. Herrero, and J. E. Suárez. 1998. The site-specific recombination system of Lactobacillus species bacteriophage A2 integrates in Gram-positive and Gram-negative bacteria. Virology. 250:185-193.
- Arfin, S.M., and H.E. Umbarger. 1969. The metabolism of valine and isoleucine in Escherichia coli. XVII. The role of induction in the derepression of acetohydroxy acid isomeroreductase. Biochem. Biophys. Res. Commun. 37:902-908.
- Auvray, F., M. Coddeville, P. Ritzenthaler, and L. Dupont. 1997. Plasmid integration in a wide range of bacteria mediated by integrase of Lactobacillus delbrueckii bacteriophage mv4. J. Bacteriol. 179:1837-1845.
- Bassit, N., C-Y. Boquien, D. Picque, and G. Corrieu. 1993. Effect of initial oxigen concentration on diacetyl and acetoin production by Lactococcus lactis subsp. lactis biovar diaetylactis. Appl. Environ. Microbiol. 59:1893-1897.
- Benson, K. H., J. –J. Godon, P. Renault, H. G. Griffin, and M. J. Gasson. 1996. Effect of ilvBN-encoded a-acetolactate synthase expression on diacetyl production in Lactococcus lactis. Appl. Microbiol. Biotechnol. 45:107-111.
- Bhowmik, T., L. Fernandez, and JL. Steele. 1993. Gene replacement in Lactobacillus helveticus. J. Bacteriol. 175:6341-6344.
- Champomier Vergès M.C., M. Zuñiga, F. Morel-Deville, G. Pérez-Martínez, M. Zagorec, and S.D. Ehrlich. 1999. Relationships between arginine degradation, pH and survival in Lactobacillus sakei. FEMS Microbiol. Lett.180:297-304.
- Dauneau, P., and G. Pérez-Martínez. 1997. Fractional factorial and multiple linear regression to optimise extraction of volatiles from a Lactobacillus plantarum bacterial suspension using purge and trap. J. Chromatogr. A 775:225-230.
- de Vos, W. M. 1999. Safe and suitable system for food-grade fermentations by genetically modified lactic acid bacteria. Int. Dairy J. 9:3-10.
- de Vos, W. M., and E. E.Vaughan. 1994. Genetics of lactose utilization in lactic acid bacteria. FEMS Microbiol. Rev. 15:217-237.
- Dickely, F., D. Nilsson, E. B. Hansen, and E. Johansen. 1995. Isolation of Lactococcus lactis non suppresors and construction of food-grade cloning vector. Mol. Microbiol. 15:839-847.
- Doi, E., D. Shibata, and T. Matoba. 1981. Modified colorimetric ninhydrin methods for peptidase assay. Anal. Biochem 118:173-184.
- Gasson, M.J. 1983. Plasmid complements of Streptococcus lactis NCDO712 and other lactic streptococci after protoplast-induced curing. J. Bacteriol. 154:1-9.
- Godon, J. -J., M. -C. Chopin, and S. D. Ehrlich. 1992. Branched-chain amino acid biosynthesis genes in Lactococcus lactis subsp. lactis. J. Bacteriol. 174:6580-6589.
- Gosalbes, M. J., V. Monedero, C.-A. Alpert, and G. Pérez-Martínez. 1997. Establishing a model to study the regulation of the lactose operon in Lactobacillus casei. FEMS Microbiol. Lett. 148:83-89.
- Gosalbes, M. J., V. Monedero, and G. Pérez-Martínez. 1999. Elements involved in catabolite repression and substrate induction of lactose operon in Lactobacillus casei. J. Bacteriol. 181:3928-3934.
- Goupil-Feuillerat, N., M. Cocaign-Bousquet, J. –J. Godon, S. D. Ehrlich, and P. Renault. 1997. Dual role of a-acetolactate decarboxylase in Lactococcus lactis subsp. lactis. J. Bacteriol. 179:6285-6293.
- Goupil, N., G. Corthier, S. D. Ehrlich, and P. Renault. 1996. Imbalance of leucine flux in Lactococcus lactis and its use for the isolation of diacetyl-overproducing strains. Appl. Environ. Microbiol. 62:2636-2640.
- Hols, P., T. Ferain, D. Garmyn, N. Bernard, and J. Delcour. 1994. Use of homologous expression-secretion signals and vector-free stable chromosomal integration in engineering of Lactobacillus plantarum for a-amylase and levanase expression. Appl. Environ. Microbiol. 60:1401-1413.
- Leenhouts, K., A. Bolhuis, G. Venema, and J. Kok. 1998. Construction of a food-grade multicopy integration system in Lactococcus lactis. Appl. Microbiol. Biotehnol. 49:417-423.
- Leloup, L., S. D. Ehrlich, M. Zagorec, and F. Morel-Deville. 1997. Single crossing-over integration in the Lactobacillus sake chromosome and insertional inactivation of the pts and the lacI genes. Appl. Environ. Microbiol. 63:2117-2123.
- Lin, M.-Y., S. Harlander, and D. Savaiano. 1999. Construction of an integrative food-grade cloning vector for Lactobacillus acidophilus. Appl. Microbiol. Biotechnol. 45:484-489.
- Maassen, C. B. M., J. D. Laman, M. J. H. den Bak-Glashouwer, F. J. Tielen, J. C. P. A. van Holten-Neelen, L. Hoogteijling, C. Antonissen, R. J. Leer, P. H. Pouwels, W. J. A. Boersma, and D. M.Shaw. 1999. Instruments for oral disease-intervention strategies: recombinant Lactobacillus casei expressing tetanus toxin fragment C for vaccination or myelin proteins for oral tolerance induction in multiple sclerosis. Vaccine 17:2117-2128.
- MacCormick, C. A., H. G. Griffin, and M. J. Gasson. 1995. Construction of a food-grade host/vector system for Lactococcus lactis based on lactose operon. FEMS Microbiol. Lett. 127:105-109.
- Martin M. C., J. C. Alonso, J. E. Suárez, and M. A. Alvarez. 2000. Generation of food-grade recombinant lactic acid bacterium strains by site-specific recombination. Appl. Environ. Microbiol. 66: 2599-2604.
- Mollet, B., J. Knol, B. Poolman, O. Marciset, and M. Delley. 1993. Directed genomic integration, gene remplacement, and integrative gene expression in Streptococcus thermophilus. J. Bacteriol. 175:4315-4324.
- Monedero, V., M. J. Gosalbes, and G. Pérez-Martínez. 1997. Catabolite repression in Lactobacillus casei ATCC 393 is mediated by CcpA. J. Bacteriol. 179:6657-6664.
- Platteeuw, C., I. van Alen-Boerrigter, S. van Schalkwijk, and W. M. de Vos. 1996. Food-grade cloning and expression systems for Lactococcus lactis. Appl. Environ. Microbiol. 62:1008-1013.
- Platteeuw, C., J. Hugenholtz, M. Starrenburg, I. van Alen-Boerrigter, and W. M. de Vos. 1995. Metabolic engineering of Lactococcus lactis: Influence of the overproduction of a-acetolactate synthase in strains deficient in lactate dehydrogenase as a function of culture conditions. Appl. Environ. Microbiol. 61:3967-3971.
- Platteeuw, C., G. Simons and W. M. de Vos. 1994. Use of the Escherichia coli b-glucuronidase (gusA) gene as a reporter gene for analyzing promoters in lactic acid bacteria. Appl. Environ. Microbiol. 60: 587-593.
- Posno, M., P. T. Heuvelmans, M. J. van Giezen, B. C. Lokman, R. J. Leer, and P. H. Pouwels. 1991. Complementation of the inability of Lactobacillus strains to utilize D-xylose with D-xylose catabolism-encoding genes of Lactobacillus pentosus. Appl. Environ. Microbiol. 57:2764-2766.
- Porter, E. V., and B. M. Chassy. 1988. Nucleotide sequence of the b-D-phospho-galactosidase gene of Lactobacillus casei: comparison to analogous pbg genes of other Gram-positive organisms. Gene 62:263-276.
- Pouwels, P. H., and R. J. Leer. 1993. Genetics of lactobacilli: plasmids and gene expression. Antoine van Leeuwenhoek 64:85-107.
- Pouwels, P. H., R. J. Leer, and W. J. A. Boersma. 1996. The potencial of Lactobacillus as a carrier for oral immunization: Development and preliminary characterization of vector systems for targeted delivery of antigens. J. Biotechnol. 44:183-192.
- Richelieu, M., U. Houlberg, and J. C. Nielsen. 1997. Determination of a-acetolactic acid and volatile compounds by headspace gas chromatografy. J. Dairy Sci. 80:1918-1925.
- Rutberg, B. 1997. Antitermination of transcription of catabolic operons. Mol. Microbiol. 23:413-421.
- Ruyter, P. G. G. A., O. Kuipers, and W. M. de Vos. 1996. Controlled gene expression systems for Lactococcus lactis with the food–grade inducer nisin. Appl. Environ. Microbiol. 62:3662-366.
- Sambrook, J., E. F. Fritsch, and T. Maniatis. 1989. Molecular Cloning: a Laboratory Manual. 2nd ed. Cold Spring Harbor Laboratory, Cold Spring Harbor, NY.
- Scheirlinck, T., J. Mahillon, H. Joos, P. Dahese, and F. Michiels. 1989. Integration and expression of a-amylase and endoglucanase genes in the Lactobacillus plantarum chromosome. Appl. Environ. Microbiol. 55:2130-2137.
- Stentz, R., and M. Zagorec. 1999. Ribose utilization in Lactobacillus sakei: analysis of the regulation of rbs operon and putative involvement of a new transporter. J. Mol. Microbiol. Biotechnol. 1:165-173.
- Stülke, J., M. Arnaud, G. Rapoport, and I. Martin-Verstraete. 1998. PRD-a protein domain involved in PTS-dependent induction and carbon catabolite repression of catabolic operons in bacteria. Mol. Microbiol. 28:865-874.
- Swindell, S. R., K. H. Benson, H. G. Griffin, P. Renault, S. D. Ehrlich, and M. J. Gasson. 1996. Genetic manipulation of the pathway for diacetyl metabolism in Lactococcus lactis. Appl. Environ. Microbiol. 62:2641-2643.
- Veyrat, A., V. Monedero, and G. Pérez-Martínez. 1994. Glucose transport by the phosphoenolpyruvate: mannose phosphotransferase system in Lactobacillus casei ATCC 393 and its role in carbon catabolite repression. Microbiology 140:1141-1149.
- Vogel, R. F., and M. Ehrmann. 1996. Genetics of lactobacilli in foods fermentations. Biotehnol. Annual Rev. 2:123-150.
- Wang, T–T., and B. H. Lee. 1997. Plasmids in Lacatobacillus. Criticals Rev. in Biotechnol. 17:227-272.
- Yebra, M. J., A. Veyrat, M. A. Santos, and G. Pérez-Martínez. 2000. Genetics of L-sorbose transport and metabolism in Lactobacillus casei. J. Bacteriol. 182:155-163.
- Zuñiga M., M.C. Champomier-Vergès, M. Zagorec, and G. Pérez-Martínez. 1998. Structural and functional analysis of the arginine deiminase gene cluster of Lactobacillus sake. J. Bacteriol. 180:4154-4159.

