Kerstin Mahr*, Carlos David Esteban*, Wolfgang Hillen, Fritz Titgemeyer and Gaspar Pérez-Martínez. * K.M and C.D.E. contributed equally to this work.
Published in Journal of Molecular Microbiology and Biotechnology (2002) 4 (5):27-32.
Pubmed link: http://www.ncbi.nlm.nih.gov/pubmed/12432959. Full text:
ABSTRACT :
In low-GC Gram-positive bacteria, carbon catabolite repression (CCR) is exerted by transcriptional regulation through a protein complex consisting of catabolite control protein CcpA and serine phosphorylated phosphocarrier protein HPr (HPr-ser-P). We investigated the interaction between these components of Lactobacillus casei and Bacillus megaterium. CcpA of L. casei could not complement a B. megaterium ccpA mutant strain, whereas it was found to be functional in Bacillus subtilis. To explore the nature of the non-complementing phenotype, we overproduced and purified CcpA and HPr of L. casei for in vitro analyses. Electrophoretic mobility shift assays revealed a failure in CCR signal transduction at the level of protein-protein interaction between L. casei CcpA and B. megaterium HPr-ser-P, while binding of CcpA to the B. megaterium target site was intact. We established a method based on surface plasmon resonance that allowed a quantitative analysis of CcpA/HPr-ser-P interactions. Calculation of the apparent dissociation constants revealed that the interaction of L. casei CcpA with B. megaterium HPr-ser-P was fivefold weaker than with its own HPr-ser-P suggesting that the reduced affinity was responsible for the non-complementing phenotype.
INTRODUCTION:
In low-GC Gram-positive bacteria, carbon catabolite repression (CCR) is exerted by energetically favorable carbon sources via the histidine-containing phosphocarrier protein HPr of the phosphoenolpyruvate-sugar phosphotransferase system (PTS) and the central regulator catabolite control protein CcpA (for a review see Hueck and Hillen, 1995; Stülke and Hillen, 1999). Upon growth on preferred sources of carbon, HPr is phosphorylated at a regulatory residue serine 46 (HPr-ser-P) by the metabolite-activated ATP-dependent HPr kinase/phosphatase HPrK (Reizer et al., 1998; Kravanja et al., 1999). HPr-serS-P is capable of interacting with CcpA promoting the specific binding to DNA target sequences termed catabolite responsive elements (cre), which are located upstream or within catabolic genes and operons (Deutscher et al., 1995; Miwa et al., 2000). This results in a repression of gene expression at the transcriptional level as has been demonstrated for a wide variety of catabolite-controlled systems including the xylose utilization operon xylAB of Bacillus megaterium (Rygus and Hillen, 1992). CcpA has been identified in bacilli, streptococci, staphylococci, and some industrially relevant lactic acid bacteria, such as Lactococcus lactis (Luesink et al) and Lactobacillus casei (Henkin et al., 1991; Egeter and Brückner, 1996; Monedero et al., 1997; Luesink et al., 1998; Simpson and Russell, 1998; Schick et al., 1999; Mahr et al., 2000). CcpA proteins were found to comprise a subfamily within the LacI/GalR family of bacterial repressors with some of the CcpA-specific residues eventually forming a continuous patch on the protein surface, which is thought to represent the binding surface of HPr-ser-P (Jones et al., 1997; Kraus et al., 1998). In addition, a mutational analysis of CcpA of B. megaterium identified amino acid positions critical for its function in CCR (Kraus and Hillen, 1997; Kraus et al., 1998; Küster et al., 1999).
To elucidate the function of CcpA in the dairy starter bacterium L. casei, we studied in vivo CCR complementation of CCR of the xylAB system of B. megaterium by L. casei CcpA in vivo. Surprisingly, we found that CcpA of L. casei was not able to confer CCR to a B. megaterium ccpA mutant, although similar heterologous cross complementation had been demonstrated earlier, including in vivo complementation of a B. subtilis ccpA mutant with L. casei ccpA (Davison et al., 1995; Egeter and Brückner, 1996; Monedero et al., 1997; Schick et al., 1999). In order to find the reason for this non-complementing phenotype, we established in vitro systems that allowed to monitor the involved protein-protein and protein-DNA interactions of L. casei and B. megaterium components. The results suggest that L. casei CcpA could not confer CCR in B. megaterium due to a reduced affinity to B. megaterium HPr-serS-P.
RESULTS:
CcpA of L. casei Cannot Complement a B. megaterium ccpA Mutant
For the in vivo complementation of a of B. megaterium ccpA mutant with ccpA of L. casei, we chose plasmid pGCCPA, which carries the L. casei ccpA gene under the control of the vegetative Gram-positive promoter SPO2 and contains a ribosome binding site and a start codon adapted to B. subtilis (Monedero et al., 1997). This construct which was derived from plasmid pGAL9 was shown to partially complement CCR of the gluconate operon gntRK of B. subtilis GM1225 (ccpA mutant) (Monedero et al., 1997). Since the plasmid-encoded erythromycin gene was not functional in B. megaterium a neomycin resistance cassette (neo) of pWH1509K was cloned in plasmids pGCCPA and pGAL9. This yielded plasmids pWH153 (ccpALca) and pWH152 (control), respectively, which were transformed into B. megaterium WH353 DccpA.
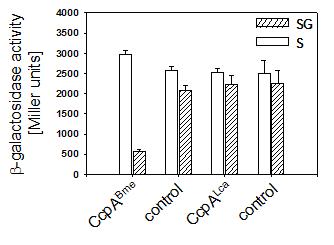
To study the ability of L. casei CcpA to confer CCR to the B. megaterium ccpA deletion mutant, we measured b-galactosidase activities of the chromosomal xylA::lacZ fusion. The xylAB promoter in this strain responds to CCR only, which is because thedue to a deletion of the xylose repressor gene was deleted rendering the system expressed in the absence of xylose. As depicted in Figure 1, B. megaterium WH353([pWH2040)], carrying ccpABme, showed a 5.2 fold reduction of b-galactosidase activity in the presence of glucose, while WH353(pWH1509K; control) was defective in CCR. Enzyme activities of strains WH353([pWH153)], carrying ccpALca, and WH353(pWH152; control) were similar under repressing conditions leading to the conclusion that CcpA of L. casei was not promoting CCR of the xyl operon in B. megaterium.
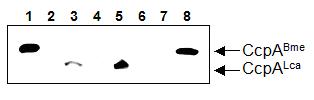
To assure that ccpA of L. casei was heterologously expressed in the B. megaterium ccpA mutant, we performed a Western blot experiment using polyclonal antibodies raised against CcpA of B. megaterium (Küster et al., 1996). CcpA could be detected in L. casei wild-type extracts, but not in the isogenic L. casei ccpA mutant (Figure 2, lanes 3 and 4). A signal was also obtained in B. megaterium WH353(pWH153, ccpALca) corresponding to the L. casei CcpA protein (lane 5). Such a band was not present in extracts of the control strain WH353(pWH152) (lane 6). CcpA of B. megaterium was readily detectable in WH353(pWH2040, ccpABme) cell extract, whereas the control extract WH353(pWH1509K) showed no CcpA signal (lanes 8 and 7). Hence, although L. casei CccpA could not complement the ccpA deletion mutant of B. megaterium, it could be shown that it wasto be indeed being expressed in B. megaterium.
Overproduction and Purification of CcpA and HPr of L. casei
In order to find an explanation for this non-complementing phenotype, we examined the in vitro interaction of L. casei CcpA with the B. megaterium target sequence crexyl in the presence and absence of B. megaterium or L. casei HPr-serS-P. For this purpose, overproduction plasmids for CcpA and HPr of L. casei were constructed and purification protocols for both proteins were established. CcpA and HPr could be purified to homogeneity following the procedure described in Materials and Methods yielding approximately 20 mg and 12 mg pure protein per liter of Escherichia coli culture, respectively. Phosphorylation of HPr of L. casei at serine 46 was achieved upon incubation of the purified protein with HPr kinase/phosphatase of B. subtilis and subsequent separation of proteins by anion exchange chromatography.
In vitro Analysis of the cre/CcpA/HPr-serS-P Complexes
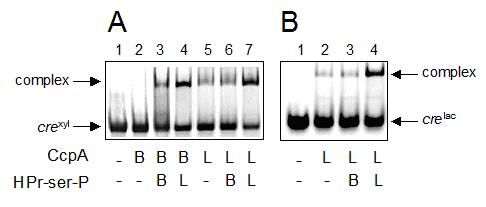
The interactions of purified components were examined by electrophoretic mobility shift assays. It could be observed (Fig. 4A) that CcpA of B. megaterium formed a complex with crexyl only in the presence of HPr-serS-PBme( (Figure 3A, lane 3). CcpA of L. casei already exhibited some binding to crexyl in the absence of HPr-serS-P under these experimental conditions (lane 5). As anticipated, a much more efficient binding of L. casei CcpA to crexyl was found when L. casei HPr-ser-P was added to the reaction, but interestingly not when HPr-ser-P of B. megaterium was used (lanes 7 and 6, respectively). This observation could be confirmed in a mobility shift assay performed using L. casei CcpA and the homologous cre sequence of the L. casei lac operon (crelac). Again, binding of L. casei CcpA to crelac was much more efficient in the presence of L. casei HPr-serS-P than with HPr-ser-P of B. megaterium (Figure 3B, lanes 4 and 3). Furthermore, there was no reciprocity in this interaction defect, since binding of B. megaterium CcpA to crexyl could be greatly enhanced by the addition of HPr-ser-P of L. casei (Figure 3A, lane 4).
Determination of Equilibrium Parameters for CcpA/HPr-serS-P Interactions
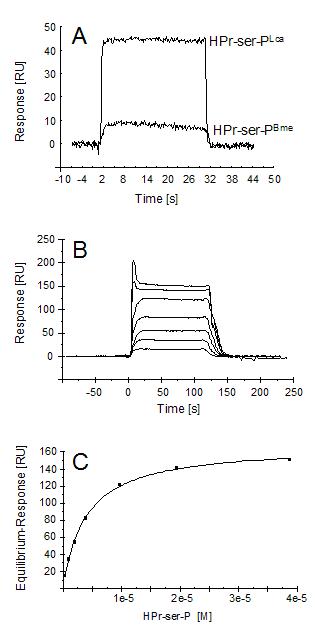
To quantify the differences in these protein-protein interactions, they were analyzed in real-time using surface plasmon resonance. L. casei CcpA was immobilizzed on a CM5 sensor chip via amino coupling. When equimolar concentrations of HPr-serS-P of L. casei or B. megaterium were applied at non-saturating CcpA-binding conditions, L. casei HPr-ser-P gave a sixfold higher resonance response signalsix fold higher, indicating that its binding ability to L. casei CcpA was correspondingly sixfold greater than that of B. megaterium HPr-ser-P (Figure 4A). Apparent equilibrium dissociation constants of the complexes of L. casei CcpA with either L. casei, B. megaterium, or B. subtilis HPr-ser-P were subsequently elucidated and found to be 3.9 x 10-6 M, 2.0 x 10-5 M, and 2.0 x 10-5 M, respectively, showing that HPr-ser-PBme and his-tagged HPr-ser-PBsu had a very similar affinity for CcpA of L. casei (Figure 4B and 4C). This affinity value was 5.1 fold lower than that of L. casei CcpA for its own HPr-ser-P, which approximately coincided with the data from resonance response signals obtained before (Figure 4A).
DISCUSSION:
This study was prompted by the initial observation that L. casei CcpA could not mediate CCR in B. megaterium, although its synthesis was readily detectable. The finding contrasted with published data, since, until date, functional complementation of CcpA between different Gram-positive bacteria had apparently been always been successful (Davison et al., 1995; Egeter and Brückner, 1996; Monedero et al., 1997; Schick et al., 1999). The lack of complementation was considered an interesting ground to set up comparative experiments that would eventually help to understand protein-protein and DNA-protein interactions taking place during the final stages of the CCR signal transduction process.
In the ternary complex cre/CcpA/HPr-ser-P, CcpA must interact with two partners and, as a consequence, the non-complementing phenotype observed could be due to a failure in an effective cre/CcpA or CcpA/HPr-ser-P interaction. Therefore, the binding ability of purified L. casei CcpA to cre sites in the presence and absence of HPr-ser-P proteins was compared by electrophoretic mobility shift assays. L. casei CcpA could efficiently recognize crexyl,Bme and crelac,Lca in the presence of its own HPr-ser-P, but not by the addition of B. megaterium HPr-ser-P. This indicated that the lack of complementation was due to a failure in the interaction of L. casei CcpA with the endogenous HPr-ser-P.
This experiment further showed that L. casei CcpA could bind cre already without corepressor, which would be a discrepancy to the in vivo results. However, this effect was considered unspecific, since L. casei CcpA exhibited some binding to the non-specific DNA included in the assay. This was monitored by Coomassie Brilliant Blue-staining of the gel shown in Figure 3A, where a major fraction of L. casei CcpA was detectable at the height of the non-specific DNA (data not shown). In contrast, purified CcpA of B. megaterium did not show unspecific binding of DNA, indicating that the binding abilities between two CcpAs cannot be directly compared with this method. Another result to discuss is that B. megaterium CcpA showed a weaker interaction with its own HPr-ser-P compared to its binding to L. casei HPr-ser-P. This shows that heterologous cross communication between two components might not always yield reciprocal results.
To determine the affinities between L. casei CcpA and different HPr-ser-P species in a more quantitative way, we chose the method of the surface plasmon resonance to monitor protein-protein interactions in real-time. This sensitive technique allowed to estimate the apparent equilibrium dissociation constants of interactions between these protein pairs, which have up to date not been described for this system (Karlsson and Falt, 1997). Using purified CcpA and HPr-ser-P of L. casei the apparent KD of the homologous interaction was 3.9 x 10-6 M. This is in good agreement with an apparent KD of 7.0 ± 2.9 x 10-6 M determined by fluorescence titration for the CcpA-homologue PurR of E. coli to its effector guanine (Xu et al., 1998). The apparent KD value obtained for the interaction of L. casei CcpA with B. megaterium HPr-ser-P was fivefold lower compared to that of both L. casei proteins.
We anticipated that HPr-ser-P of B. subtilis would exhibit a higher binding affinity to L. casei CcpA since in vivo complementation was reported (Monedero et al., 1997). Surprisingly, the KD was the same as determined for the pair CcpALca/HPr-ser-PBme. Obviously, similar heterologous cross communication was sufficient for complementation of CCR in B. subtilis, while it did not work in B. megaterium. It should be noted that CcpA of L. casei complemented only partially (about 40%) CCR of the B. subtilis gnt target system (Monedero et al., 1997). Thus, other parameters such as protein expression levels, the choice of the reporter system, and the presence of a second HPr-ser-P-like protein (Crh-ser-P) could have accounted for this difference (Galinier et al., 1997).
As mentioned before, functional interaction between HPr-ser-P and CcpA belonging to different species has been demonstrated, but has not been studied at the molecular level. The here established method to quantitatively determine CcpA/HPr-ser-P interactions may provide a useful tool to investigate CcpA and HPr protein variants in a combinatorial manner.
EXPERIMENTAL PROCEDURES:
Bacterial Strains, Plasmids and Growth Conditions
The strains and plasmids used in this work are listed in Table 1. L. casei cells were grown in MRS medium (Difco) at 37ºC under static conditions. E. coli was grown with shaking at 37ºC in Luria-Bertani (LB) medium. B. megaterium cells were grown in LB medium and M9 mineral medium supplemented with 25 mM succinate at 28ºC with shaking. Glucose at a concentration of 10 mM was added when repressing growth conditions were required. Plating of bacteria was performed on the same media solidified with 1.5% agar. When appropriate, the concentrations of antibiotics used were 100 µg/ml of ampicillin or 30 µg/ml of kanamycin for E. coli and 4 µg/ml of neomycin for B. megaterium.
Construction of L. casei ccpA Expression Vectors
A neomycin resistance cassette (neo) of pWH1509K was cloned in plasmids pGCCPA and pGAL9 using standard procedures and E. coli DH5a as host (Sambrook et al., 1989). The fragment containing the neo cassette was obtained by XbaI restriction of pWH1509K and inserted into pGCCPA and pGAL9 digested with the same restriction enzyme yielding plasmids pWH153 and pWH152, respectively.
Enzyme Assays
Quantification of b-galactosidase activity in B. megaterium cells was carried out as described by Hueck et al. (Hueck et al., 1995). Cells were grown in M9 mineral medium supplemented with 25 mM succinate (non-repressing growth conditions) or with 25 mM succinate plus 10 mM glucose (repressing growth conditions). Experiments were repeated in triplicate.
Western Blot Analysis
B. megaterium strains derived from WH353(DccpA) transformed with plasmid pWH153 carrying L. casei ccpA (ccpALca), pWH152 (control), pWH2040 carrying B. megaterium ccpA (ccpABme), and pWH1509K (control), were grown in LB medium supplemented with neomycin. L. casei strains ATCC393 (wild type) and BL71 (ccpA mutant) were grown in MRS medium (Difco). Cells were harvested at an OD600 of 0.5 by centrifugation and cell extracts were subsequently prepared as described previously (Monedero et al., 1997; Mahr et al., 2000). Proteins of cell extracts were separated by sodium dodecyl sulfate-polyacrylamide gel electrophoresis on a 7.5% polyacrylamide gel and transferred to a polyvinylidene difluoride membrane (Fluorotrans) by electroblotting. CcpA was detected with a rabbit polyclonal antiserum raised against CcpA of B. megaterium (Küster et al., 1996). CcpA antibodies on the polyvinylidene difluoride membrane were visualized using the ECL Western blot analysis system (Amersham).
Table 1. Strains and plasmids used in this study.
strain/plasmid | genotype/description | reference |
B. megaterium WH353 | DccpA gdh2f(xylA1-spoVG-lacZ) DxylR | Küster et al., 1999 |
L. casei BL23 | ATCC 373 [pLZ15–] | B. Chassy (University of Illinois, Urbana) |
L. casei BL71 | L. casei BL23 ccpA mutant | V. Monedero et al., 1997 |
E. coli DH5a | general cloning host | Sambrook et al., 1989 |
E. coli BL21 | protein overproduction strain (DE3, pLysS) | Studier, 1991 |
pGal9 | erm | Monedero et al., 1997 |
pGCCPA | erm, pGal9 containing ccpA gene (L. casei) | V. Monedero et al., 1997 |
pWH1509K | neo | Rygus and Hillen, 1992 |
pWH2040 | neo, pWH1509K containing ccpA gene (B. megaterium) | Hueck et al., 1995 |
pWH152 | pGAL9 plus neo from pWH1509K | This work |
pWH153 | pGCCPA plus neo from pWH1509K | This work |
pET3c | bla | Studier et al., 1990 |
pET3c-ccpA | pET3c containing ccpA of L. casei | This work |
pET3c-ptsH | pET3c containing ptsH of L. casei | This work |
Construction of CcpA and HPr Overproduction Vectors
Plasmid pET3c-ccpA for the overproduction of L. casei CcpA was constructed by amplifying the ccpA gene from plasmid pUCCPA (Monedero, unpublished) with oligonucleotides CCPA1 (GGAGAAAATCATATGGAAAAGC) and CCPA2 (AAAAGGATCCATTATTTCGTTG) introducing NdeI and BamHI restriction sites (underlined), respectively. PCR was performed with Pfu DNA polymerase. The restricted amplification product was cloned in the expression vector pET3c digested with the same restriction enzymes yielding pET3c-ccpA. Plasmid pET3c-ptsH for overproduction of HPr of L. casei was obtained by a similar procedure. The ptsH gene (encoding HPr) of L. casei was amplified from chromosomal DNA with oligonucleotides PTSH1 (CAGATCACATATGGAAAAACGCG) and PTSH2 (AAATGTGGATCCATTATTCAGCC). All constructs were verified by DNA sequencing.
Protein Overproduction and Purification
For protein purification, plasmids pET3c-ccpA and pET3c-ptsH were transformed in E. coli BL21(DE3)(pLysS) (Studier, 1991). Overexpression of proteins was achieved by isopropyl thio-b-D-galactoside (IPTG) induction as described (Parche et al., 1999). For the purification of CcpA, one liter of E. coli culture was grown in the presence of 1 mM IPTG for 3 h. Cells were harvested by centrifugation and washed with buffer A (20 mM Tris-HCl (pH 8.0), 2 mM dithiothreitol). The pellet was resuspended in 5 ml of buffer A and crude extracts were prepared by sonification at 45 W 5 x 30 s (Labsonic U, Braun) and subsequent removal of cell debris by centrifugation. The cleared cell extract was used for an anion exchange chromatography (HQ/M 16/100; Poros) using a gradient from 0 to 1 M NaCl. Fractions containing CcpA were pooled, concentrated, and proteins were further separated by gel filtration (Superdex G75; Pharmacia) using a buffer containing 20 mM Tris-HCl (pH 8.0), 2 mM dithiothreitol, and 200 mM NaCl, giving a homogeneous pure CcpA preparation. Purification of heat-stable HPr was achieved following a similar procedure with the exception that the cleared cell extract was boiled for 10 min and centrifuged to remove the majority of cell proteins. The supernatant was subjected to anion exchange chromatography as described above yielding fractions containing homogeneous pure HPr.
In vitro Phosphorylation of HPr at Serine 46
HPr of L. casei, B. megaterium, or B. subtilis was phosphorylated at serine 46 with HPr kinase/phosphatase from B. subtilis expressed from p4813 as previously described (Kraus et al., 1998; Reizer et al., 1998).
Electrophoretic Mobility Shift Assays
Two small DNA fragments that contained characterized cre sites were used in these experiments. A synthetic 48 bp double-stranded oligonucleotide containing the cre sequence of the B. megaterium xylA gene (position +105 to +152 of xylA) (crexyl) (5 µM) or a 26 bp double-stranded oligonucleotide comprising the cre sequence of the L. casei lac operon (position -23 to -48 of lacT) (5 µM) were mixed with purified CcpA (10 µM) of B. megaterium or L. casei with or without 10 µM HPr-ser-P of B. megaterium or L. casei in a total volume of 10 µl. Samples were incubated in the presence of 2 µg unspecific DNA (linearized plasmid pWH802) at 37ºC in a buffer containing 100 mM Tris-HCl (pH 7.5) and 1 mM EDTA for 10 min before loading on a non-denaturing 5% polyacrylamide gel. Gels were run with a buffer containing 100 mM Tris-HCl (pH 8.8) and 1 mM EDTA at 70 V for 45 min. DNA was visualized by ethidium bromide staining.
Surface Plasmon Resonance Experiments
The interaction of L. casei CcpA with HPr-ser-P from L. casei, B. megaterium, or B. subtilis was analyzed by surface plasmon resonance with a BiaCore X biosensor (Pharmacia Biosensor AB). CcpA was immobilized on a CM5 sensor chip (Pharmacia Biosensor AB) by amino coupling to 1.5 to 1.7 ng/mm2 following instructions supplied by the manufacturer. The difference in CcpA binding between L. casei and B. megaterium HPr-ser-P was determined by subsequently flowing these phosphorylated effectors over the sensor chip in HBS-EP buffer (BiaCore AB) at a flow rate of 100 ml/min. To determine equilibrium constants, a set of sensorgrams was collected at a flow rate of 5 ml/min using various concentrations of L. casei or B. megaterium HPr-ser-P or his-tagged HPr-ser-P from B. subtilis. Apparent equilibrium dissociation constants of the complexes were estimated by plotting the equilibrium response against HPr-ser-P concentrations and fitted to the model Req = KA x C x Rmax / KA x C x n+1, where Req is the response in equilibrium, KA the apparent association constant (inverse of KD), C the concentration of HPr-ser-P, Rmax the maximal response (when the chip reaches saturation) and n a sterical impediment correction factor. This fitting allowed the calculation of the apparent KD of protein-protein interactions. Sensorgrams were analyzed using the BIAevaluation 3.1 software (BiaCore AB).
ACKNOWLEDGEMENTS:
K. M. and C. D. E. contributed equally to this work.
We thank Elke Küster-Schöck for gifts of protein and antibodies and for helpful discussions. We are grateful to Matthias Merzbacher, Andrea Wagner, Lwin Mar Aung-Hilbrich, and Marco Diel for gifts of DNA and protein.
This work was financed by the EU project BIO4-CT96-0380 and by funds of the Spanish CICyT (Interministerial Commission for Science and Technology) (Ref. ALI 98-0714). C. D. E. was the recipient of a fellowship from the Spanish government. K. M. was supported by a grant from the Graduiertenkolleg Kontrolle der RNA-Synthese.
References:
Davison, S.P., Santangelo, J.D., Reid, S.J. and Woods, D.R. 1995. A Clostridium acetobutylicum regulator gene (regA) affecting amylase production in Bacillus subtilis. Microbiology 141: 989-996.
Deutscher, J., Küster, E., Bergstedt, U., Charrier, V. and Hillen, W. 1995. Protein kinase-dependent HPr/CcpA interaction links glycolytic activity to carbon catabolite repression in Gram-positive bacteria. Mol. Microbiol. 15: 1049-1053.
Egeter, O. and Brückner, R. 1996. Catabolite repression mediated by the catabolite control protein CcpA in Staphylococcus xylosus. Mol. Microbiol. 21: 739-749.
Galinier, A., Haiech, J., Kilhoffer, M.C., Jaquinod, M., Stülke, J., Deutscher, J. and Martin-Verstraete, I. 1997. The Bacillus subtilis crh gene encodes a HPr-like protein involved in carbon catabolite repression. Proc Natl Acad Sci U S A 94: 8439-8444.
Henkin, T.M., Grundy, F.J., Nicholson, W.L. and Chambliss, G.H. 1991. Catabolite repression of a-amylase gene expression in Bacillus subtilis involves a trans-acting gene product homologous to the Escherichia coli lacI and galR repressors. Mol. Microbiol. 5: 575-584.
Hueck, C.J. and Hillen, W. 1995. Catabolite repression in Bacillus subtilis: a global regulatory mechanism for the Gram-positive bacteria? Mol. Microbiol. 15: 395-401.
Hueck, C.J., Kraus, A., Schmiedel, D. and Hillen, W. 1995. Cloning, expression and functional analyses of the catabolite control protein CcpA from Bacillus megaterium. Mol. Microbiol. 16: 855-864.
Jones, B.E., Dossonnet, V., Küster, E., Hillen, W., Deutscher, J. and Klevit, R.E. 1997. Binding of the catabolite repressor protein CcpA to its DNA target is regulated by phosphorylation of its corepressor HPr. J Biol Chem 272: 26530-26535.
Karlsson, R. and Falt, A. 1997. Experimental design for kinetic analysis of protein-protein interactions with surface plasmon resonance biosensors. J Immunol Methods 200: 121-133.
Kraus, A. and Hillen, W. 1997. Analysis of CcpA mutations defective in carbon catabolite repression in Bacillus megaterium. FEMS Microbiol Lett 153: 221-226.
Kraus, A., Küster, E., Wagner, A., Hoffmann, K. and Hillen, W. 1998. Identification of a co-repressor binding site in catabolite control protein CcpA. Mol Microbiol 30: 955-963.
Kravanja, M., Engelmann, R., Dossonnet, V., Bluggel, M., Meyer, H.E., Frank, R., Galinier, A., Deutscher, J., Schnell, N. and Hengstenberg, W. 1999. The hprK gene of Enterococcus faecalis encodes a novel bifunctional enzyme: the HPr kinase/phosphatase. Mol Microbiol 31: 59-66.
Küster, E., Hilbich, T., Dahl, M.K. and Hillen, W. 1999. Mutations in catabolite control protein CcpA separating growth effects from catabolite repression. J. Bacteriol. 181: 4125-4128.
Küster, E., Luesink, E.J., de Vos, W.M. and Hillen, W. 1996. Immunological crossreactivity to catabolite control protein CcpA from Bacillus megaterium is found in many Gram-positive bacteria. FEMS Microbiol. Lett. 139: 109-115.
Luesink, E.J., van Herpen, R.E., Grossiord, B.P., Kuipers, O.P. and de Vos, W.M. 1998. Transcriptional activation of the glycolytic las operon and catabolite repression of the gal operon in Lactococcus lactis are mediated by the catabolite control protein CcpA. Mol Microbiol 30: 789-798.
Mahr, K., Hillen, W. and Titgemeyer, F. 2000. Carbon catabolite repression in Lactobacillus pentosus: analysis of the ccpA region. Appl. Environ. Microbiol. 66: 277-283.
Miwa, Y., Nakata, A., Ogiwara, A., Yamamoto, M. and Fujita, Y. 2000. Evaluation and characterization of catabolite-responsive elements (cre) of Bacillus subtilis. Nucleic Acids Res 28: 1206-1210.
Monedero, V., Gosalbes, M. and Perez-Martinez, G. 1997. Catabolite repression in Lactobacillus casei ATCC 393 is mediated by CcpA. J. Bacteriol. 179: 6657-6664.
Parche, S., Schmid, R. and Titgemeyer, F. 1999. The phosphotransferase system (PTS) of Streptomyces coelicolor: identification and biochemical analysis of a histidine phosphocarrier protein HPr encoded by the gene ptsH. Eur J Biochem 265: 308-317.
Reizer, J., Hoischen, C., Titgemeyer, F., Rivolta, C., Rabus, R., Stülke, J., Karamata, D., Saier, M.H.J. and Hillen, W. 1998. A novel protein kinase that controls carbon catabolite repression in bacteria. Mol. Microbiol. 27: 1157-1169.
Rygus, T. and Hillen, W. 1992. Catabolite repression of the xyl operon in Bacillus megaterium. J Bacteriol 174: 3049-3055.
Sambrook, J., Fritsch, E.F. and Maniatis, T. 1989 Molecular cloning: a laboratory manual. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, N. Y.
Schick, J., Weber, B., Klein, J.R. and Henrich, B. 1999. PepR1, a CcpA-like transcription regulator of Lactobacillus delbrückii ssp. lactis. Microbiology 145: 3147-3154.
Simpson, C.L. and Russell, R.R. 1998. Identification of a homolog of CcpA catabolite repressor protein in Streptococcus mutans. Infect Immun 66: 2085-2092.
Studier, F.W. 1991. Use of bacteriophage T7 lysozyme to improve an inducible T7 expression system. J Mol Biol 219: 37-44.
Studier, F.W., Rosenberg, A.H., Dunn, J.J. and Dubendorff, J.W. 1990. Use of T7 RNA polymerase to direct expression of cloned genes. Methods Enzymol 185: 60-89.
Stülke, J. and Hillen, W. 1999. Carbon catabolite repression in bacteria. Curr Opin Microbiol 2: 195-201.
Xu, H., Moraitis, M., Reedstrom, R.J. and Matthews, K.S. 1998. Kinetic and thermodynamic studies of purine repressor binding to corepressor and operator DNA. J Biol Chem 273: 8958-8964.

